Join us as a Masters or PhD student: Chemical reactions in lattice polymer models
Required experience in the following methods:
Statistical physics, molecular dynamics, monte carlo, lattice simulations
Work with:
- Dr. Tyler Harmon | Tel: +49 (351) 4658 1216 | Email: TylerHarmon@ipfdd.de
- Prof. Dr. Jens-Uwe Sommer | Tel: +49 (351) 4658 750 | Email: Sommer@ipfdd.de
where: Leibniz Institute for Polymer Research Dresden (Leibniz-Institut für Polymerforschung (IPF) Dresden), Germany
Project description:
Biomolecular condensates are a newly defined class of organelle in the cell. In contrast to the classical picture of a cellular organelle, there is no membrane separating the inside from outside. Instead, they form through the physics of proteins phase separating into droplets similar to oil demixing from water. One of their primary functions is expected to be as reaction chambers where enzymes and substrates are localized into volumes separated from the rest of the cell. Much of the work incorporating chemical reactions into droplets has been through studying reaction diffusion equations or solving Cahn-Hilliard equations. These techniques are all mean field and could miss important physics that becomes apparent at the polymer scale.
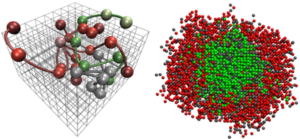
In this project we will build upon a lattice polymer model to develop and incorporate chemical reactions into this framework. Lattice polymer models provide an important intermediate level of coarse-graining which has been used successfully for studying how proteins phase separate. Understanding chemically active systems requires access to systems even larger than what is needed to study the thermodynamics, making lattice polymer models especially important. The larger scale view of where this project fits into is understanding how polymer architecture can impact the chemical reactions. Some examples are: (i) Does the heteropolymer nature that is inherent in proteins enhance droplets’ ability to act as chemical reactors? (ii) Are there cases where caging effects can be beneficial? (iii) What are the limits where reaction-diffusion equations break down?
 The key tasks for the project are
The key tasks for the project are
- Add an additional move to the simulation engine that changes one class of bead to different class of bead.
- Characterize how to relate the monte carlo moves to the dynamics of a real system.
- Run simulations with different architectures of polymers to examine how the macroscopic scale reaction rates are impacted.
For more information: https://www.ipfdd.de/de/forschung/institut-theorie-der-polymere/students/topics-for-master-theses/chemical-reactions-in-lattice-polymer-models/
Current news by our research groups
Anthony Hyman Group,Agnes Toth-Petroczy Group
CD-Code is now published in Nature Methods
 CD-CODE is now published in Nature Methods. It is a “living database” that we designed for fast…
CD-CODE is now published in Nature Methods. It is a “living database” that we designed for fast…
Alf Honigmann Group,Anthony Hyman Group,Marcus Jahnel Group,Simon Alberti Group
A role for RNA in Stress Granules assembly
Stress granules are membraneless compartments formed by phase separation of specific molecules upon exposure to cellular stress such as oxidative stress, heat shock, or osmotic stress. The Alberti, Jahnel, Honigmann, and Hyman labs published a study in cell highlighting the role of RNA in the…
Filament formation by the translation factor eIF2B regulates protein synthesis in starved cells
Aminoacyl-tRNA synthetases (aaRSs), the enzymes responsible for coupling tRNAs to their cognate amino acids, minimize translational errors by intrinsic hydrolytic editing. Here, we compared norvaline (Nva), a linear amino acid not coded for protein synthesis, to the proteinogenic, branched valine…
Anthony Hyman Group,Moritz Kreysing Group,Simon Alberti Group
Condensation regulates translation
New insights into the influence of Ded1p condensation on translation comes from the Hyman, Alberti and Kreysing labs. The study published in Cell is entitled "Condensation of Ded1p Promotes a Translational Switch from Housekeeping to Stress Protein Production".
Graphical abstract: